3 The Experiment
Uropathogenic Escherichia coli (UPEC) employs a complex set of molecular factors that allow it to evade host defences, resist antibiotic treatment, and persist in the urinary tract, ultimately facilitating pathogenesis.
To remain in the urinary tract UPEC employs adherence factors, molecules that act to help UPEC adhere to the tissue of the channel itself, or to material in that channel, such as catheter tubes. If we can intervene to reduce this adherence, we can shorten the dwell time of the microbe and reduce the impact of disease.
3.1 Experimental overview
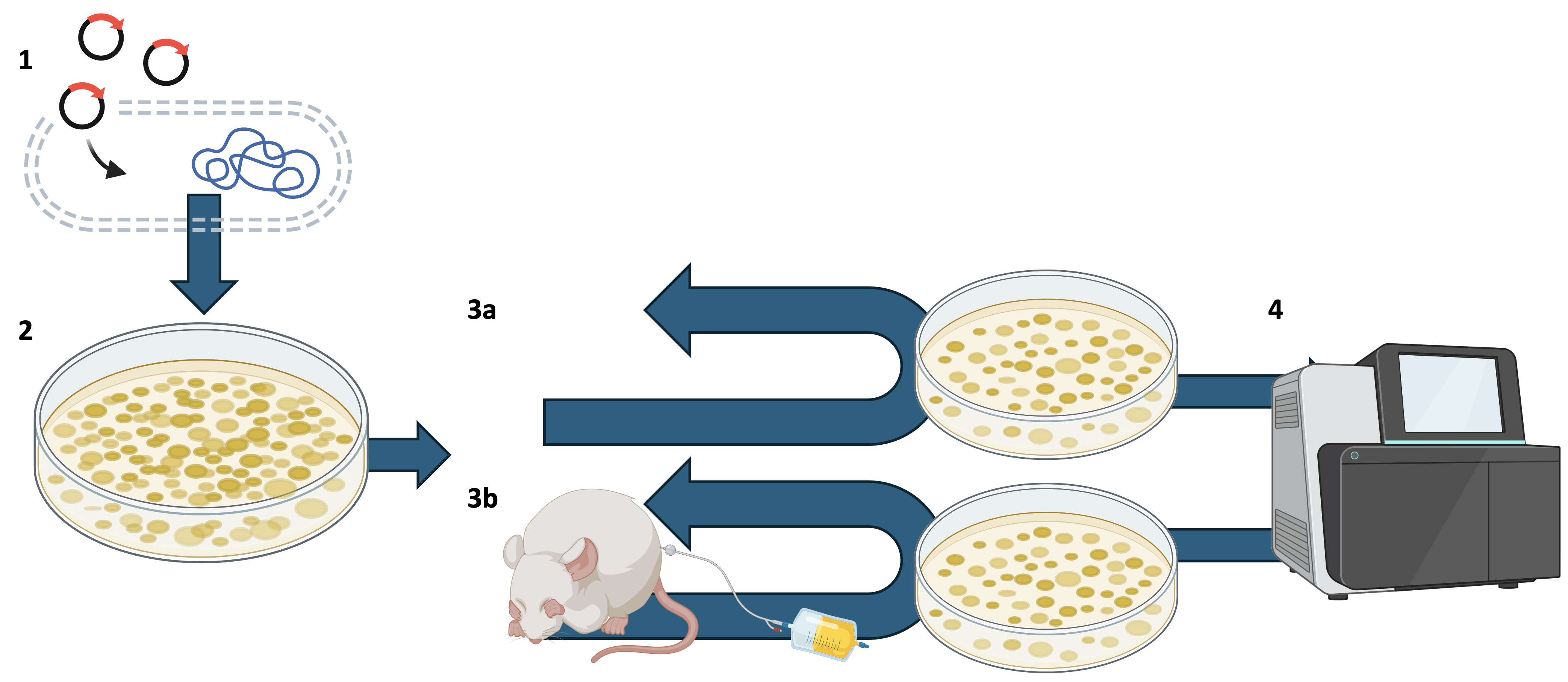
The overall plan of the experiment is sketched in Figure 3.1.
A bacterial artificial chromosome (BAC) library is constructed for an E. coli UPEC isolate.
A bacterial artificial chromosome is a DNA construct that introduces a large genomic region (typically 150-350kbp) from a genome of interest into a “neutral” background genome. In this experiment, genomic regions from UPEC are introduced into a “neutral” background E. coli (DH10B). This allows us to investigate the effect of genes from UPEC when expressed by an otherwise non-pathogenic E. coli.
The UPEC genome was partially digested with Hind III and cloned into a pV41 vector. This was used to generate three pools of 384 BAC clones to facilitate replication.
The BAC library is then passaged through a mouse model (i.e. a catheterised mouse) to emulate introduction to the human catheterised urinary tract. After passage and washing of the extracted catheter tube, the tube was used as inoculum for a further round of incubation to culture adherent bacteria.
Passage is the serial, iterated growth of bacteria under a particular environment or series of environments. This approach is usually taken to subject the bacteria to a specific selection pressure so that we can observe how it evolves in response to that pressure.
This is a relatively common approach to investigating bacterial virulence factors.
Mouse passage was allowed to proceed for 18h. Subsequent culturing was performed in LB media at 37 degC for 18h between passages.
A parallel control passage, identical except that bacteria were cultured in LB for 18h rather than in the mouse model, was also performed.
Three rounds of passage were performed.
After passage, the total gDNA from the three control and three mouse system cultures is sequenced. This produces read counts corresponding to three kinds of genes:
- genes present only in the background DH10B
- genes present in both DH10B and UPEC
- genes present only in UPEC and carried through passage
3.2 Expected interpretation
We are most interested in the genes that are present only in UPEC, and which form part of the BAC library.
We expect that the serial passage process in the mouse model arm of the experiment enriches our cultures for bacteria that adhere better to the urinary tract and/or catheter material. As individual clones contain only fragments of the UPEC genome, those clones that don’t contain adherence factors are expected to be selected out of the mixture, leaving only clones with UPEC genes that promote adherence.
It is also possible that some UPEC genes confer an advantage for survival in the passage process itself, rather than an advantage specific to adherence. Both the control and mouse model arms of the experiment are assumed to enrich for these genes equally.
By statistical analysis of the read counts, we aim to identify UPEC-specific genes that are overrepresented in the final mixtures that have been through mouse passage, but that are not overrepresented in the control passage through culture alone. These are the genes we will take forward as candidates for involvement in catheter adherence.
You can see how well this worked, if at all, in the next section.